std_statfieldtrip missing interaction (Robert -> can be implemented with differences)
Description
Dear Arnaud,
I wanted to pointed out that it seems like the computation of the interaction for an ANOVA design (example below) relying on fieldtrip statistics is not possible through the std_stat function, is it done on purpose?
#### Steps to Reproduce
- [First Step: Random data]
data = { rand(50,10) rand(50,10);rand(50,12) rand(50,12)}; % 2 groups, 2 conditions = mixed ANOVA
- [Second Step: Fieldtrip permutation statistics]
[Res.pcond, Res.pgroup, Res.pinter,Res.statscond, Res.statsgroup, Res.statsinter] = ...
std_stat(data,'condstats','on','groupstats','on','paired',{'off' 'on'},'fieldtripnaccu',5000,'fieldtripmethod','montecarlo',...
'fieldtripmcorrect','max','fieldtripalpha',0.05,'mode','fieldtrip');
- [Third Step: Outputs]
As can be seen below the third cells in
pinterandstatsinterare empty...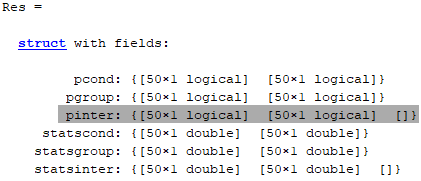
#### Actual behavior: In line 201-202 in std_stat you assign to `pinter` and `statsinter` variables an empty cell of 1x3 (i.e. from what I understand for main effects + interaction)
pinter = cell(1,3);
statsinter = cell(1,3);
Then in lines 223-228 you compute the first main effect:
% main statistics
if ng > 1
[F, df, pval] = statcondfieldtrip(newdata, 'paired', opt.paired{1}, params{:});
pinter{1} = applymask(F, opt.fieldtrip);
statsinter{1} = squeeze(F.stat);
end
And then lines 252-257 the second main effect:
% main statistics
if nc > 1
[F, df, pval] = statcondfieldtrip(newdata, 'paired', opt.paired{1}, params{:});
pinter{2} = applymask(F, opt.fieldtrip);
statsinter{2} = squeeze(F.stat);
end
But you never assign anything to the tird cell pinter{3} and statsinter{3}.
Is it because the computation of an interaction is not possible using Fieldtrip-lite?
Thanks and best wishes,
Corentin
#### Versions
| OS version | Windows 10 Enterprise build 17763 |
| Matlab version | Matlab R2019b |
| EEGLAB version | EEGLAB v.2020 (development version from GitHub) |
Yes, I do not believe this is implemented in Fieldtrip (has it changed?) so there is no way to do it.
I will have a close look in Fieldtrip and let you know, many thanks for your reply.
OK let me know. I think I look a couple of months ago and it still was not there. I could be mistaken though.
Dear Arnaud,
I opened an issue on Fieldtrip GitHub and just got reply.
According to this tutorial, it is actually possible to compute the interaction term using cluster-based permutation tests for factorial designs which don't have more than 1 between-subjects factor.
Hence, in my case of a 2x2 mixed design, it should be possible.
Let me know what you think about it.
Best,
Corentin
OK, I will see if I can change the code in statcondfieldtrip to do that.
Many thanks Arnaud!
Hi Corentin,
I have revisited this issue in depth. The functions are just not compatible when more than one independent variable is used.
ft_statfun_indepsamplesF ft_statfun_depsamplesFmultivariate
I have looked in detail if I was calling the functions properly. The code is simply not compatible when cfg.ivar contains more than one index. Copying @robertoostenveld just in case. However, it should be possible to use the Matlab statistics toolbox which has come a long way. Or you can use LIMO which allows using cluster correction for multiple comparisons for complex designs.
Arno
The concept of a randomization test on a multi-factorial design - where you are usually interested in a (non-linear) interaction - is explained here http://www.fieldtriptoolbox.org/faq/how_can_i_test_an_interaction_effect_using_cluster-based_permutation_tests/. In short, you should test for differences in the differences.
Thank you Robert. I will look it up.