Error handling for processes
We would like NF to support metadata tracking and routing; this would for example enable a process to inform another process that an action was not taken or a stage not passed, this could trigger other tasks. I include two examples from our current pipeline below; perhaps not a perfect match, but they informed my initial thinking about this.
process irods {
tag "${samplename}"
input:
val samplename from sample_list.flatMap{ it.readLines() }
output:
set val(samplename), file('*.cram') optional true into ch_cram_files
file('*.lostcause.txt') optional true into ch_lostcause_irods
script:
"""
if bash -euo pipefail irods.sh -t ${params.studyid} -s ${samplename}; then
true
else
stat=\$?
tag='UNKNOWN'
if [[ \$stat == 64 ]]; then tag='nofiles'; fi
echo -e "${samplename}\\tirods\\t\$tag" > ${samplename}.lostcause.txt
fi
"""
}
# later ...
ch_star_accept = Channel.create()
ch_star_reject = Channel.create()
star_aligned
.choice(ch_star_accept, ch_star_reject)
{ namelogsbams -> check_log(namelogsbams[1]) ? 0 : 1 }
ch_star_accept
.map { name, logs, bams -> [name, bams] }
.into { ch_featurecounts; ch_indexbam }
ch_star_reject
.map { it -> "${it[0]}\tSTAR\tlowmapping\n" }
.mix(ch_lostcause_irods)
.set { ch_lostcause }
Additional comment: the IRODS errors I am tracking above may be transient; if I rerun the pipeline hours or a day later this stage may then succeed. However, in this set-up, Nextflow will never retry execution upon resuming the pipeline with -resume. If I were to change the code and have non-zero exit status in the failing else branch (where tag=UNKNOWN), then my tracking information would not be sent forward. So at the moment my custom meta-data tracking system is not fully functional; either I can have -resume working for the irods process or I can have tracking, but I can't have both.
I think what it could be done is to add some new process event handlers e.g. onSuccess, onError, onComplete and use to track specific tasks events and metadata. For example:
process irods {
tag "${samplename}"
input:
val samplename from sample_list.flatMap{ it.readLines() }
output:
set val(samplename), file('*.cram') optional true into ch_cram_files
script:
"""
irods.sh -t ${params.studyid} -s ${samplename}
"""
onError:
if( task.exitStatus==64 )
ch_lostcause_irods << [process: task.process, failed:true, sampleName: sampleName]
}
The idea is to allow process event handlers to access the same process evaluation context including task runtime info and input/output files metadata.
That looks good. In onError, would the code still have access to files produced by the script section? I don't have a scenario, but perhaps worth considering?
Just looking at our rnaseq pipeline, I have this tracking in two places, but need it in at least two more places, and generally I would probably use this type of code in the majority of my processes. (see also previous comment).
would the code still have access to files produced by the script section?
It should be possible to access to output files metadata ie. file names, size, etc.
Following from: https://gitter.im/nextflow-io/nextflow?at=5bce925b384492366182345b @pditommaso
Process event handlers would be really great - another use case below:
I usually pass input and execution metadata around a pipeline without passing around additional files. This goes hand-in-hand with NF principles where we (mostly) don't need to keep track of file names or use them for storing information. For example a read alignment process which can work with multiple read aligners can look like this (each *meta is a map object):
process align {
tag("${idxmeta} << ${readsmeta}")
input:
set val(idxmeta), file("*"), val(readsmeta), file(r1), file(r2) from indices.combine(reads)
output:
set val(meta), file("*bam") into BAMs
script:
meta = idxmeta.clone() + readsmeta.clone()
template "${idxmeta.tool}_align.sh"
}
In this particular case it would be best to store some execution information in the meta before it is passed downstream. This could, for example, be information stored by NF in .command.trace or more generally from any file generated by the script block. It is easy enough to declare such files as output and pass them through the pipeline and parse in a downstream process, but if there are multiple files like this things will get messy.
Interesting. Yes, the idea would be to expose in the completion handler runtime and input/output files metadata. However in principle it should not allow files I/O manipulation. Do you think this would fit your use case ?
For this use case reading file content is necessary, but not writing.
Currently, modifying the example above I can declare .command.trace as the third element of output set
output:
set val(meta), file("*bam"), file('.command.trace') into BAMs
then I can parse and discard the trace file from the set before the downstream process
process downstreamProcess {
input:
set val(meta), file(sam) from alignedDatasets.map { meta, sam, trace ->
meta.'aligntrace' = trace.splitCsv( header: true, limit: 1, sep: ' ')
meta.'aligntrace'.'duration' = trace.text.tokenize('\n').last()
new Tuple(meta, sam)
}
Given NFs trace report generation this is not normally necessary, but the file in question could be a log file generated by some tool executed within the script block.
If it is possible to read/parse such files immediately after completion of the script block that could be really useful.
This is a fairly basic question I fear; with the above code, so
onError:
if( task.exitStatus==64 )
ch_lostcause_irods << [process: task.process, failed:true, sampleName: sampleName]
How can I later serialise these tuples to a file that can be controlled similar to publishDir? Is this possible in what you propose? I'm still very eager for this feature!
You could use ch_lostcause_irods as any other channel in a downstream process.
I'm a bit slow ... I want to collect all the values from the channel, stick it into a file, and ideally do 1) publish it 2) send it also to MultiQC. I found collectFile and can do 1), but don't yet see how I can do 2). (This assumes the feature as suggested above):
Channel
.from( ['irods', 'lung', 'not ready'],
['cram', 'gut', 'lowreads'],
['STAR', 'kidney', 'lowmapping'] )
.set { ch_lostcause }
ch_lostcause
.map { it.join("\t") + "\n" }
.collectFile(name: 'lostcause.txt', storeDir: "$baseDir/results")
Assuming input as above with this feature I'd like to achieve both 1) and 2). Perhaps it is already possible now, and I missed the solution, but I thought it worth documenting this scenario.
Just to remark that in Simone Coughlan's talk at Nextflow 'Reproducible In silico Genomics' it seemed that she is doing similar accounting tasks in Nextflow code: https://github.com/coughls/nextflow-example-scripts . Edit: tagging @coughls .
I'm thinking that this approach is still too programmatic oriented. At the end the real need here is to capture tasks metadata and save to a file (or eventually to a database, but this could be a sub-scenario).
It think NF should provide a more declarative approach ie. the process should have a meta directive that allows you to declare the metadata attributes you to track. Then the system can collect all these info and save automatically to a file.
Allright, it sounds like a principled approach. I'll stay tuned to see what happens. This could enable me to create a report as sketched above; but if it is something that happens after workflow completion then I won't be able to send it to MultiQC. I'll try to see if there is something simple that allows me to do this in pure Nextflow; my main current issue is that I cannot have normal resumption and error tracking at the same time. As I see it there are two different issues:
- General error/metadata tracking in Nextflow pipelines, for which your proposed
metais perfect. - More bespoke tracking for cases of particular interest, where the user will appreciate an application-specific report, such as a. data not ready b. not enough data c. not enough data mapped.
In case (2), there really is interest in a limited set of processes within the pipeline, and it would be nice to manage this in the pipeline by e.g. sending it to something like MultiQC. Do you think the distinction between (1) and (2) is a valid one, or would you say the proposed meta will take care of both?
I would really love to have a better integration with MultiQC. How these info would populate the MultiQC report? Your idea is to create a custom file and add it as an input to the MultiQC step?
This is working at the moment, as custom content. This is example data that I create using the lostcause channels:
# plot_type: 'table'
# section_name: 'Lost samples'
Sample Process Message
ImmHet6564407 cram lowreads
ImmHet6564790 cram lowreads
ImmHet6565818 cram lowreads
ImmHet6565945 cram lowreads
The comment lines are to instruct MultiQC. This results in this multiqc section:
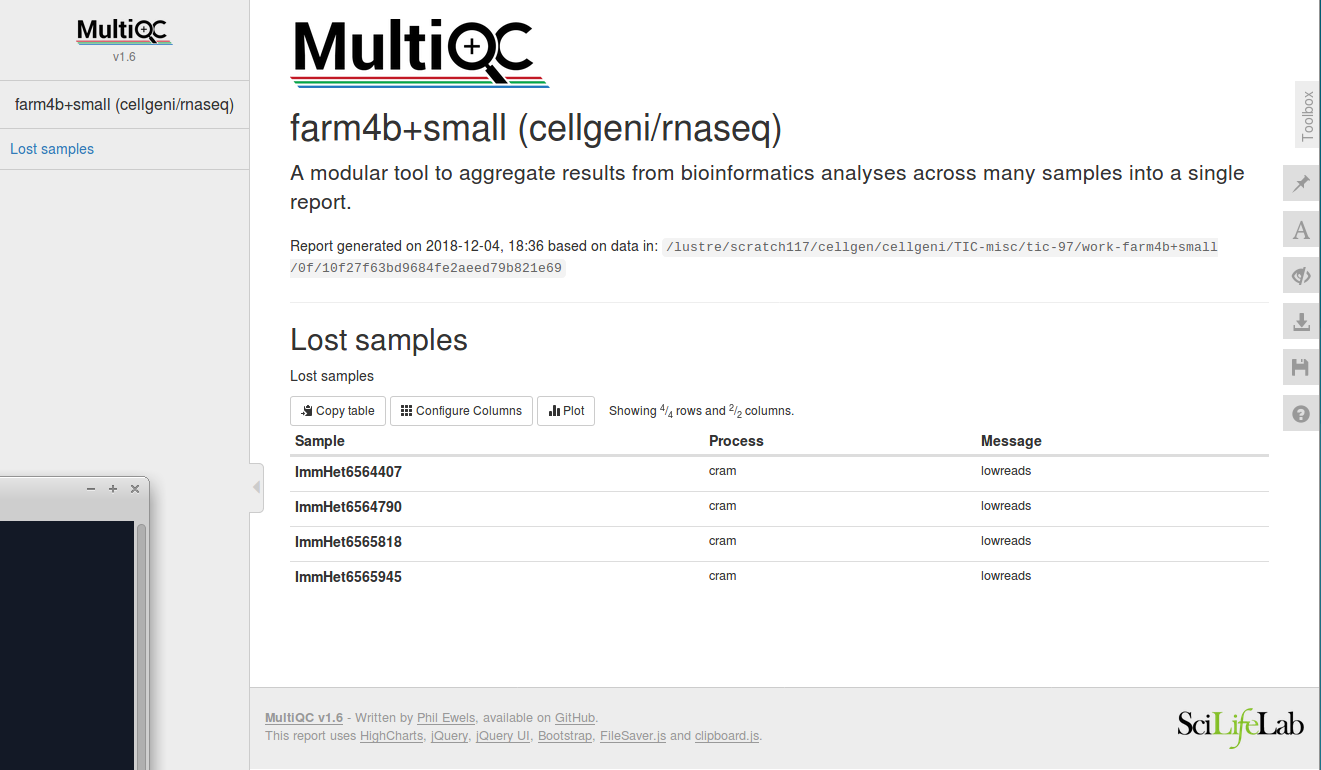 The work directory and command:
The work directory and command:
farm,0f/10f27f63bd9684fe2aeed79b821e69, ls
farm4b+small_multiqc.html farm4b+small_multiqc_data lostcause
farm,0f/10f27f63bd9684fe2aeed79b821e69, cat .command.sh
#!/bin/bash -euo pipefail
multiqc . -f --title "farm4b+small (cellgeni/rnaseq)" --filename "farm4b+small_multiqc.html" -m custom_content -m featureCounts -m star -m fastqc
The way I see it now (but happy to learn more) is that MultiQC and nextflow are both very flexible and supply a lot of plumbing options, so I think nothing multiqc-specific is needed in Nextflow. But some generic support in Nextflow could still be useful. I've implemented this for both STAR and hisat2 now, in what seems the best idiom I can come up with:
ch_hisat2_accept = Channel.create()
ch_hisat2_reject = Channel.create()
ch_hisat2_aligned
.choice(ch_hisat2_accept, ch_hisat2_reject)
{ namelogbams -> hisat2_filter(namelogbams[1]) ? 0 : 1 }
ch_hisat2_accept
.map { name, log, bam -> [name, bam] }
.set { ch_hisat2_bam }
ch_hisat2_reject
.map { it -> "${it[0]}\thisat2\tlowmapping\n" }
.mix(ch_lostcause_irods, ch_lostcause_cram, ch_lostcause_star)
.set { ch_lostcause }
Another pattern I've used twice is this:
.... // irods process.
output:
set val(samplename), file('*.cram') optional true into ch_cram_files
file('*.lostcause.txt') optional true into ch_lostcause_irods
... // samtools crams to fastq process (this tests for a minimum number of reads in the script section).
output:
set val(samplename), file("${samplename}_?.fastq.gz") optional true into ch_fastqs_irods
file('*.lostcause.txt') optional true into ch_lostcause_cram
...
This one (two channels, optional file names) is actually pretty sweet and short; but for processes where the error is potentially transient (as in the irods process) it stops making the process being activated again under -resume. However, this is perhaps an exceptional case; maybe it's actually not that high among my priorities.
A short update: After the above I've concluded that what I'd like most is to be able to catch unforeseen process failures, e.g. a program crashing or not producing an output unexpectedly, and then manage that failure as a first-class citizen in the dataflow layer, e.g. include this in the same 'lost cause' report. I would like to be able to do this without instrumenting every single process. I'm thinking a generic 'onError' with a closure that has a little bit of context (for example the tag and exit status, perhaps the input names and STDERR output). Perhaps a process could optionally set up some extra custom metadata that can be part of that context.
I am also interested in this, as a method to allow external programs to parse the output of Nextflow pipelines with higher-level logic instead of having to do something like hard-code in the publishDir locations and file naming patterns to look for, which might change during development. Would be great to have some record of all the meta-data surround each task execution, especially paths to output and publishDir files, along with all the other evaluated directives included in each task (e.g. things described here).
Hi @micans @stevekm
The above would be quite useful. What is the current status with the above? I think the best way forward would be to organize a potential PR, with @pditommaso involved.
Thanks
A further thought (experiment). A very general solution in this problem space could be the option to specify that a channel can still be activated when a process fails. This would in one fell swoop allow me to do anything that I want with that error using the full power of Nextflow. The default behaviour would be as it is now; on a task error all its output channels are closed to the task. One or more output channels could be specified to remain open if an error occurs, perhaps by specifying a closure that for example receives the exit code and perhaps some more context.
Importantly, the task would still be resumed with -resume. I like this idea as it is very generic and flexible and does not commit to any framework, hopefully it is not at odds with fundamental design principles.
This would allow various logging methods; for example logging everything, or logging only faulty processes if a channel is for example equipped both with onError: open and its inputs as optional: True.
My current idea is to allow the implementation of event handlers on each process scope. For example having
process foo {
input: ..
output: ..
"your_command .."
}
Then it should be possible to do:
foo.onError {
// logic here to handle the error code
// `task` implicit variable allow the access to runtime metada, inputs and outputs
}
foo.onComplete {
// etc
}
A generic process should be accessible as
process.onError { .. etc }
It should even be possible to pipe the result of event handles to a channel, e.g.
foo.onError { /* logic */ } | error_channel_name
This sounds great Paolo. And this:
It should even be possible to pipe the result of event handles to a channel, e.g.
foo.onError { /* logic */ } | error_channel_name
Is exactly what I'm hoping for, as it allows a generic user report to be constructed with normal processes, publishDir et cetera, and it will be easy for example to embed it in a multiQC report. The task implicit variable would have the metadata and exit code so this creates a lot of power for bespoke reporting.
Hi @pditommaso
My current idea is to allow the implementation of event handlers on each process scope.
I think this makes a great deal of sense, thanks! This should address quite a few concerns.
Do you have an outline how we the community could best start on this PR?
Also, I very much agree with @micans here:
this ... is exactly what I'm hoping for, as it allows a generic user report to be constructed with normal processes, publishDir et cetera, and it will be easy for example to embed it in a multiQC report. The task implicit variable would have the metadata and exit code so this creates a lot of power for bespoke reporting.
Let me know how to best help out---this feature would be invaluable!
YES! I'd love to see something like this. I'll echo the call for errors becoming first class citizens in the dataflow so that I can write processes to handle the errors later in my pipeline. I really want to be able to do something along these lines:
nextflow.preview.dsl=2
process foo {
input:
val(x)
output:
file("${x}.txt")
script:
"""
# error prone script to create ${x}.txt
"""
onError:
{ /* some closure to handle the error */ }
}
foo()
other_process(foo.out)
error_collection_process(foo.err)
So, the key ideas are:
- Allow each process to specify its own
onErrorclosure. - The closure has access to error metadata as well as input data. Saying that process
foofailed is useful, but saying that processfoofailed with input${x}is really useful. - Put the result of the error handler closure into a channel. I'm imagining that every process would have a default
emptyerror channel and then only if a handler is provided would the error channel be populated.
To me getting the error into a channel is the most important part so that I can integrate those errors downstream and produce the output I'm required to.
Hello, there are quite a few ideas on how to approach this. How would you feel @pditommaso about the following small new feature:
- an output channel can declare e.g.
onFail: true(similar tooptional: true). - if a process fails, only output channels with
onFail: trueare sent to. - on resume, a failed process will still be re-tried.
This looks to me like a smaller feature, rather than a grand over-arching framework; Perhaps both such a smaller feature and the grand framework could coexist; I'm hoping something like this is small and dedicated and hopefully not too intrusive to implement .
I just created a repo demonstrating a hacked up way to handle errors in almost the way I want things to work. https://github.com/mes5k/nf_error. The key requirement for me is to be able to get some sort of error data (including process inputs) into a channel to be used within the pipeline. In the README for the repo I've described the problems I see with my approach.
@pditommaso if you'd be willing to provide a few pointers on where one might add support for a feature like this, I'd be happy to try and put a PR together.
I still think this ticket is very much worth pursuing, and I would be happy to help out with a PR. I think we just need some internal agreement/direction upon what precisely needs to be done. :)
Re-reading this ticket, it reminds me of the approach that I demo'd here; https://github.com/stevekm/nextflow-communicator
To manually save meta data to a file in each process then read it with an external service
This would be better handled by the onComplete and onError handling described here though.
This issue has been automatically marked as stale because it has not had recent activity. It will be closed if no further activity occurs. Thank you for your contributions.