Wrong last stable nucleotide for a deletion in GRCh37 to GRCh38 conversion
Dear Authors,
Thank you for this great tool!
I have been testing CrossMap with DBSNP v151 data and found one type of conversion that doesn't look correct.
For DBSNP rs202079816: given chr22:28084772 CT>C on GRCh37 as input, CrossMap produces chr22:27688775 GT>C on GRCh38 as output, where I would expect chr22:27688775 GT>G on GRCh38.
Here the ALT seems not to be set to the last stable nucleotide (G) in GRCh38. Note that the last stable nucleotide differ between the two assemblies GRCh37 (C) and GRCh38 (G). Does this affect only deletions or also other variant types?
I used CrossMap (v0.6.1) and chain file: ftp://ftp.ensembl.org/pub/assembly_mapping/homo_sapiens/GRCh37_to_GRCh38.chain.gz
Many Thanks, Ivan
Hi Ivan, Maybe I did not fully understand the problem here. CrossMap only updates the reference allele but not the alternative allele. So here CT>C => GT>G, Why C was changed into G? Is this variant on the "-" strand? What you mean "the last stable nucleotide" ?
Hi Liguo,
Thank you for your reply and sorry for my previous imprecise description. I think, this is probably a bug in CrossMap, I have found a similar issue for 930 DBSNP variants. Below, you can find one example:
The input VCF variant, GRCh37 coordinates: #CHROM POS ID REF ALT QUAL FILTER INFO chr22 28084772 rs202079816 CT C . . RS=202079816
The output VCF variant in the CrossMap output, GRCh38 coordinates: chr22 27688775 rs202079816 GT C . . RS=202079816
Expected VCF variant, GRCh38 coordinates: chr22 27688775 rs202079816 GT G . . RS=202079816
Corresponding mapping represented by the chain file, when running highly appreciated: CrossMap.py viewchain GRCh37_to_GRCh38.chain.gz chr22 28067972 28085507 + 22 27671975 27689510 +
In the reference genome GRCh37 at position chr22:28084773, there is a T. In GRCh38 at position chr22:27688776 there is also a T.
In GRCh37 at position chr22:28084772, there is a C. In GRCh38 at position chr22:27688775 there is a G.
When running the same input variant through GATK LiftoverVcf, the variant gets rejected with: MismatchedRefAllele, which I would understand, as there is a mismatch in the reference allele (C vs G).
For deletions, according to the VCF standard, the first nucleotide in REF and ALT correspond to the nucleotide at POS in the reference genome, I would therefore expect that in the case the first nucleotide in REF is adjusted by CrossMap, then the first nucleotide in ALT should also be set to the same nucleotide (i.e. G in the example above).
To conclude, I think that this variant should either be rejected or the nucleotide in the ALT should be set to G. Where, I would prefer to still map this variant with corrected ALT. Or I am missing something and a deletion can turn into a delins?
Many Thanks, Ivan
Hi Ivan, Thank you very much for clarifying. This makes sense to me now. We will update and release a new version ASAP.
Liguo
Hi Liguo,
Thank you very much, I am looking forward to the new version. Today, I think I have found another issue: https://github.com/liguowang/CrossMap/issues/47
Best wishes, Ivan
Hi Liguo,
Thank you very much for releasing CrossMap version 0.6.3. I now run the full test with more than 600 million DBSNP v151 variants.
With CrossMap version 0.6.1, I found 930 problematic variants targeted in this issue. With the new version, I am still unfortunately getting 109 problematic variants, which is a big improvement, although the problem now seems to be for variants on the negative strand on GRCh38.
Example:
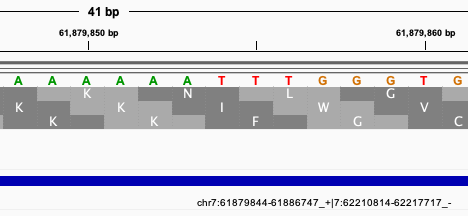
For GRCh37 input variant: chr7 61879851 rs1223781306 A AC . . .
I would expect according to DBSNP on GRCh38: chr7 62217710 rs1223781306 T TG . . .
Nevertheless, I got from CrossMap on GRCh38: chr7 62217710 . T GA . . .
The problem seems to be that an insertion turns into a delins, which does not seem to be correct. The REF and ALT should start with the REF nucleotide T at position chr7:62217710 on GRCh38. Would it be possible to fix also such variants?
Attached is a file containing all the 109 problematic variants, where INFO.GRCh37 is the input GRCh37 variant in format CHROM:POS:REF:ALT. Fields CHROM, POS, REF, ALT contain the CrossMap v0.6.3 output on GRCh38. The ID contains the expected mapping according to DBSNP on GRCh38 in format: s;RS;GENE;CHROM:POS:REF:ALT, where the ALT can contain multiple values for one RS identifier. I am also aware that not all expected DBSNP variant mappings are correct according to the chain file I am using, where I am using: ftp://ftp.ensembl.org/pub/assembly_mapping/homo_sapiens/GRCh37_to_GRCh38.chain.gz
Best wishes, Ivan CrossMap_v063_issue_44.txt
Thanks, Ivan. This is very helpful. I will look into these issues.
Hi Liguo,
I would like to ask you, whether it is planned to fix the remaining issues described here and in #47 in the near future? I plan to lift over a big set of variants, so it would be helpful to know, whether I should wait for a new version or use the current CrossMap version?
Many Thanks, Ivan
We will try to address this issue in a few weeks, sorry all of us have busy schedules. will let you know...
Many Thanks, I am looking forward to the new version
Hi Liguo,
Thank you for releasing the 0.6.4 CrossMap version.
I tested the lift over from GRCh37 to GRCh38 using ensembl chain files with ~660 million DBSNP v151 variants and the results were identical to the 0.6.3 CrossMap version.
Unfortunately, the issue described here and in #47 have not yet been fixed. I understand you are very busy, but do you have an estimate when it could be fixed?
Many Thanks, Ivan
I too am having issues where insertions become deletions. It seems when using Crossmap MAF, the tool doesnt actually look at the original reference (-) and just shoves in the asssemblies reference. This causes some insertions to become SNVs or deletions.
Hello, I've experienced issues whose origin are from these changes (I think) - that's why I'm posing here (issue should be reopened maybe?):
I've checked VCF spec and found what @ivan-mh was talking about - if we have GGT REF and G-T ALT - it isn't chr1:149032050 GT->T - it is chr1:149032049 GG->G.
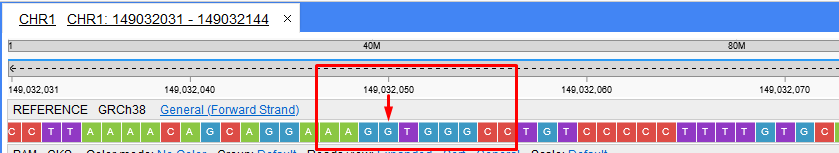
But here is the problem - before we have incorrectly stated indels:
chr1 149032050 indel.6062 GT T . PASS . GT:GQ 0/0:3
chr1 148974590 indel.6073 AG G . PASS . GT:GQ 0/0:3
chr1 148953847 indel.6081 CA A . PASS . GT:GQ 0/0:2
chr1 148953579 indel.6083 GA A . PASS . GT:GQ 0/0:3
chr1 145994150 indel.6100 C TC . PASS . GT:GQ 0/0:2
chr1 145994018 indel.6101 CAGTATA A . PASS . GT:GQ 0/0:2
chr1 145994017 indel.6102 T CT . PASS . GT:GQ 0/0:2
chr1 145892823 indel.6130 T CTCTTT . PASS . GT:GQ 0/0:3
chr1 145873621 indel.6147 TG G . PASS . GT:GQ 0/0:2
chr1 145873607 indel.6148 G TG . PASS . GT:GQ 0/0:3
chr1 145872926 1KG_1_145562155 T CCTT . PASS . GT:GQ 0/0:2
chr1 145867368 indel.6157 TG G . PASS . GT:GQ 0/0:3
chr1 145838109 indel.6181 TAG G . PASS . GT:GQ 0/0:2
chr1 145826589 indel.6187 AC C . PASS . GT:GQ 0/0:3
chr1 145736151 indel.6189 CGTAAGCACCCCAAGCA A . PASS . GT:GQ 0/0:3
chr1 145731005 indel.6192 GT T . PASS . GT:GQ 0/0:3
chr1 145686476 indel.6198 C AC . PASS . GT:GQ 0/0:6
chr1 144423146 var_1_148594506 TG G . PASS . GT:GQ 0/0:2
chr1 206016023 indel.9616 G AGGAG . PASS . GT:GQ 0/0:3
chr10 47355450 indel.12475 TG G . PASS . GT:GQ 0/0:2
chr10 47353516 indel.12476 GT T . PASS . GT:GQ 0/0:3
chr10 46010865 indel.12613 TC C . PASS . GT:GQ 0/0:2
chr11 54707629 indel.17968 G TG . PASS . GT:GQ 0/0:3
chr11 54707548 indel.17970 CT T . PASS . GT:GQ 0/0:3
chr11 54603389 indel.17972 T AT . PASS . GT:GQ 0/0:2
chr11 54603365 indel.17973 G CAG . PASS . GT:GQ 0/0:2
chr15 23030984 indel.34217 GTTGA A . PASS . GT:GQ 0/0:2
chr15 23021965 indel.34223 C GC . PASS . GT:GQ 0/0:3
chr15 23011305 indel.34227 G AG . PASS . GT:GQ 0/0:2
chr21 10592374 indel.69325 GT T . PASS . GT:GQ 0/0:1
But now we have ... I can't understand, what do we have now. Some random alleles.
chr1 149032050 indel.6062 GT C . PASS . GT:GQ 0/0:3
chr1 148974590 indel.6073 AG T . PASS . GT:GQ 0/0:3
chr1 148953847 indel.6081 CA G . PASS . GT:GQ 0/0:2
chr1 148953579 indel.6083 GA C . PASS . GT:GQ 0/0:3
chr1 145994150 indel.6100 C TG . PASS . GT:GQ 0/0:2
chr1 145994018 indel.6101 CAGTATA G . PASS . GT:GQ 0/0:2
chr1 145994017 indel.6102 T CA . PASS . GT:GQ 0/0:2
chr1 145892823 indel.6130 T CTCTTA . PASS . GT:GQ 0/0:3
chr1 145873621 indel.6147 TG A . PASS . GT:GQ 0/0:2
chr1 145873607 indel.6148 G TC . PASS . GT:GQ 0/0:3
chr1 145872926 1KG_1_145562155 T CCTA . PASS . GT:GQ 0/0:2
chr1 145867368 indel.6157 TG A . PASS . GT:GQ 0/0:3
chr1 145838109 indel.6181 TAG A . PASS . GT:GQ 0/0:2
chr1 145826589 indel.6187 AC T . PASS . GT:GQ 0/0:3
chr1 145736151 indel.6189 CGTAAGCACCCCAAGCA G . PASS . GT:GQ 0/0:3
chr1 145731005 indel.6192 GT C . PASS . GT:GQ 0/0:3
chr1 145686476 indel.6198 C AG . PASS . GT:GQ 0/0:6
chr1 144423146 var_1_148594506 TG A . PASS . GT:GQ 0/0:2
chr1 206016023 indel.9616 G AGGAC . PASS . GT:GQ 0/0:3
chr10 47355450 indel.12475 TG A . PASS . GT:GQ 0/0:2
chr10 47353516 indel.12476 GT C . PASS . GT:GQ 0/0:3
chr10 46010865 indel.12613 TC A . PASS . GT:GQ 0/0:2
chr11 54707629 indel.17968 G TC . PASS . GT:GQ 0/0:3
chr11 54707548 indel.17970 CT G . PASS . GT:GQ 0/0:3
chr11 54603389 indel.17972 T AA . PASS . GT:GQ 0/0:2
chr11 54603365 indel.17973 G CAC . PASS . GT:GQ 0/0:2
chr15 23030984 indel.34217 GTTGA C . PASS . GT:GQ 0/0:2
chr15 23021965 indel.34223 C GG . PASS . GT:GQ 0/0:3
chr15 23011305 indel.34227 G AC . PASS . GT:GQ 0/0:2
chr21 10592374 indel.69325 GT C . PASS . GT:GQ 0/0:1
Fortunately its only ~20 indels across 10000 indels in our data but this is still a problem.
Btw - I'm using https://hgdownload.cse.ucsc.edu/goldenpath/hg19/liftOver/hg19ToHg38.over.chain.gz chain - should I use ftp://ftp.ensembl.org/pub/assembly_mapping/homo_sapiens/GRCh37_to_GRCh38.chain.gz instead?
To be honest, I don't know which one (chr1:149032050 GT->T or chr1:149032049 GG->G) is correct. But for the 2nd one (chr1:149032049 GG->G) would NOT tell you which reference G is deleted. right?
According to spec - second one is correct because only it has correct ALT in POS. And how can you differentiate which one G is deleted in nature? Aren't they the same?
I agree. The 2nd one follows the VCF specification. This should be an easy fix.
In regard to CROSSMAP MAF and how it handles indels, shouldI open a separate issue?
In regard to CROSSMAP MAF and how it handles indels, shouldI open a separate issue?
We will do the same way as VCF files
@ivan-mh , Sorry for the long delay. The issue (#47) is a little bit complicated. Could you please check if this beta version fixes the problem? I hope this change will not mess up other variants (It does work for rs768155704). Thanks very much.
https://drive.google.com/file/d/16HsEz-CcxOgjuzN5-9Utx_Txn8D_vOAr/view?usp=sharing
On Mon, Aug 1, 2022 at 6:38 AM ivan-mh @.***> wrote:
Hi Liguo,
Thank you for releasing the 0.6.4 CrossMap version.
I tested the lift over from GRCh37 to GRCh38 using ensembl chain files with ~660 million DBSNP v151 variants and the results were identical to the 0.6.3 CrossMap version.
Unfortunately, the issue described here and in #47 https://github.com/liguowang/CrossMap/issues/47 have not yet been fixed. I understand you are very busy, but do you have an estimate when it could be fixed?
Many Thanks, Ivan
— Reply to this email directly, view it on GitHub https://github.com/liguowang/CrossMap/issues/44#issuecomment-1201086278, or unsubscribe https://github.com/notifications/unsubscribe-auth/ACN443RVLJKHQ5RE764JFWTVW6ZLVANCNFSM5OU7U3QQ . You are receiving this because you modified the open/close state.Message ID: @.***>
@liguowang Hello, do we have fix for this issue?
@liguowang Hello, do we have fix for this issue?
please download the code (https://drive.google.com/file/d/16HsEz-CcxOgjuzN5-9Utx_Txn8D_vOAr/view?usp=sharing) to see if the problem is solved or not. Thanks
I've installed b0 version:
#11 [ 7/10] RUN cd "/soft" && wget -q .../distr/CrossMap-0.6.5b0.tar.gz -O "/soft/CrossMap-0.6.5b0.tar.gz" && tar -xzf "/soft/CrossMap-0.6.5b0.tar.gz" && cd "/soft/CrossMap-0.6.5b0" && pip3 install --no-cache-dir .
#11 0.644 Processing /soft/CrossMap-0.6.5b0
#11 0.645 Preparing metadata (setup.py): started
#11 0.850 Preparing metadata (setup.py): finished with status 'done'
#11 0.857 Requirement already satisfied: cython>=0.17 in /usr/local/lib/python3.7/dist-packages (from CrossMap==0.6.5b0) (0.29.34)
#11 0.858 Requirement already satisfied: pysam in /usr/local/lib/python3.7/dist-packages (from CrossMap==0.6.5b0) (0.21.0)
#11 1.185 Collecting bx-python
#11 1.385 Downloading bx_python-0.9.0-cp37-cp37m-manylinux_2_17_x86_64.manylinux2014_x86_64.whl (4.3 MB)
#11 1.668 ━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━ 4.3/4.3 MB 15.4 MB/s eta 0:00:00
#11 1.829 Collecting pyBigWig
#11 1.884 Downloading pyBigWig-0.3.22-cp37-cp37m-manylinux_2_17_x86_64.manylinux2014_x86_64.whl (192 kB)
#11 1.888 ━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━ 192.5/192.5 kB 150.1 MB/s eta 0:00:00
#11 1.892 Requirement already satisfied: numpy in /usr/local/lib/python3.7/dist-packages (from bx-python->CrossMap==0.6.5b0) (1.21.6)
#11 1.896 Building wheels for collected packages: CrossMap
#11 1.897 Building wheel for CrossMap (setup.py): started
#11 2.193 Building wheel for CrossMap (setup.py): finished with status 'done'
#11 2.194 Created wheel for CrossMap: filename=CrossMap-0.6.5b0-py3-none-any.whl size=39444 sha256=8df307f6559be4273b5f56d539dd40467e21fa33595e58254756fa319683226e
#11 2.194 Stored in directory: /tmp/pip-ephem-wheel-cache-miuf2i4c/wheels/f6/e7/83/788fdaa5286f29f5194ab664a6c17dfe88f61b8dd8a70f19be
#11 2.196 Successfully built CrossMap
#11 2.241 Installing collected packages: pyBigWig, bx-python, CrossMap
#11 2.641 Successfully installed CrossMap-0.6.5b0 bx-python-0.9.0 pyBigWig-0.3.22
Nothing changed:
< chr1 149032050 indel.6062 GT T . PASS . GT:GQ 0/0:3
---
> chr1 149032050 indel.6062 GT C . PASS . GT:GQ 0/0:3
25383c25383
< chr1 148974590 indel.6073 AG G . PASS . GT:GQ 0/0:3
---
> chr1 148974590 indel.6073 AG T . PASS . GT:GQ 0/0:3
25408c25408
@liguowang Issue is not fixed - do you have any other ides how to fix it?
@liguowang Issue is not fixed - do you have any other ides how to fix it?
I will see if I have some time this evening to look into this issue.
@liguowang If you need help with fix or feedback - we can try, but we don't know where to look at now.
According to VCF specs (Section 5.1.1 Example 1), the first nucleotide of ALT should always be the same as the first nucleotide of REF. right? If this is true, the variants below are NOT valid:
chr1 149032050 indel.6062 GT T . PASS . GT:GQ 0/0:3 chr1 148974590 indel.6073 AG G . PASS . GT:GQ 0/0:3 chr1 148953847 indel.6081 CA A . PASS . GT:GQ 0/0:2 chr1 148953579 indel.6083 GA A . PASS . GT:GQ 0/0:3 ...
Yes, you are correct - these variants are incorrectly written in VCF. We have this conversation from here - chr1:149032050 GT->T should be chr1:149032049 GG->G. But at least we can understand these deletions.
In old versions deletions have last nucleotide of ALT same as the last nucleotide of REF, but correct will be first, as you pointed out.
Hello again, any chance that you'll find time to fix?
@liguowang Hello! Any news? This is important for me too. Help us, please!
@liguowang Hello! Any news? This is important for me too. Help us, please! This is not an easy fix. I just released v0.6.6, but this version still has the same issue probably. I will take a close look after vacation (August 3). Sorry for the delay.
This is not an easy fix. I just released v0.6.6, but this version still has the same issue probably. I will take a close look after vacation (August 3). Sorry for the delay.
Thank you very much!