Using lambda function in combine_vertices dictionary of cluster_graph causes memory leak
Hello, When using lambda function in the combine_vertices attribute of cluster_graph method (see code) I discovered a memory leak: after deleting the condensed graph, a section of memory is still unreachable. If no lambda function is used, the memory leak does not happen.
strongly_connected = graph.clusters(mode="strong")
condensed = strongly_connected.cluster_graph( combine_vertices=dict( name=lambda x: x, min_count='min', orig_id_graph=lambda x: x, ) )
I needed to create a graph condensation on a large graph (~10^6 vertices) several times (10000+), and by trying to do so, I caused a memory overflow (on a 32GB RAM). From the documentation, I was under the impression that this functionality is supported. I am able to surpass this by using an external dictionary, however, I thought you should know about the leak.
Thank you for an otherwise great package.
Version information: I installed python-igraph 0.9.1 via pip and I am using Python 3.8.
Thanks a lot, I'll look into this! It looks like python-igraph is accidentally holding on to a reference of a Python object that should otherwise be freed.
So I've spent a bit of time debugging the problem, but I could not reproduce it so far.
My setup was:
- python-igraph
masterbranch (but no related change since 0.9.1 so it should be the same as 0.9.1) - Python 3.9
- macOS 10.15.7
I used the following script to test things:
from igraph import Graph
from random import shuffle
num_nodes = 6400
numbers = list(range(num_nodes))
shuffle(numbers)
graph = Graph.GRG(num_nodes, 0.01)
graph.vs["name"] = ["name{}".format(i) for i in range(num_nodes)]
graph.vs["min_count"] = numbers
for i in range(200000):
strongly_connected = graph.clusters(mode="strong")
condensed = strongly_connected.cluster_graph(combine_vertices=dict(name=lambda x: x, min_count='min', orig_id_graph=lambda x: x, ))
I then used the Leaks module of Instruments.app to take snapshots of allocations every 10 seconds, but I could see no signs of increasing memory usage; see the attached screenshot.
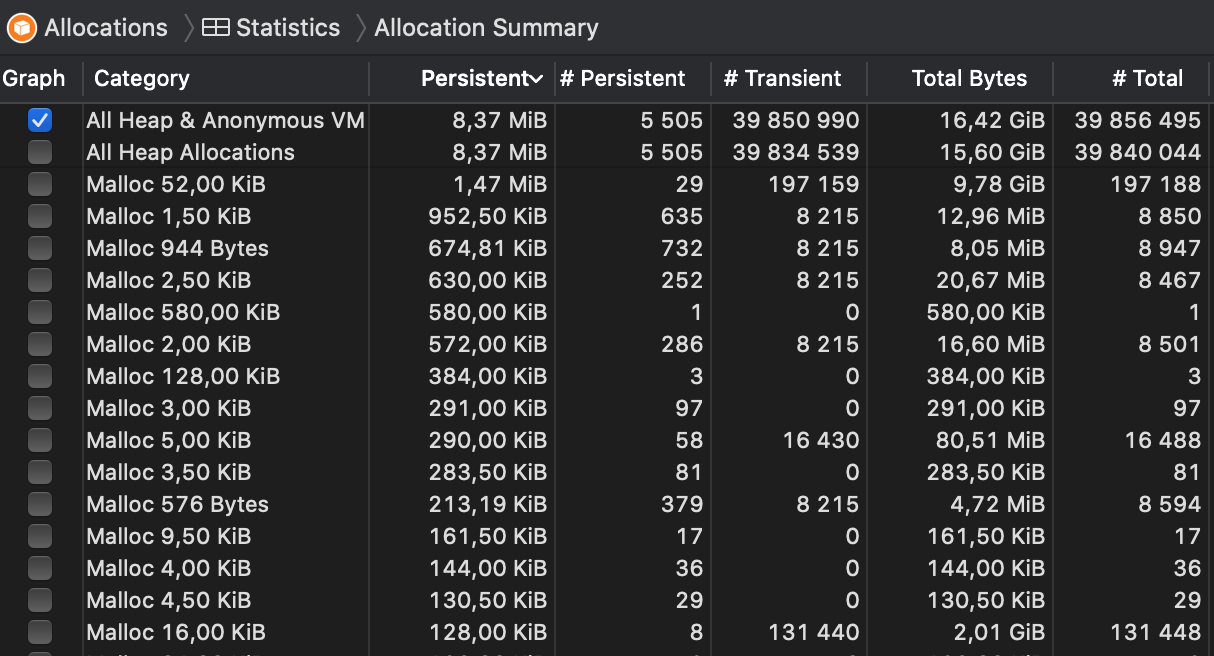
Basically, 16.42 GiB of memory was allocated during the test (which took about two minutes), but the vast majority of it was transient allocation that was then freed subsequently. The persistent allocations were floating at 8.37 MiB after the few seconds and stayed there for the whole duration.
I'll try again with larger graphs tomorrow, but I think there must be something that I'm missing in my reproduction attempts. Can you share a bit more details about what exactly you are trying to do and how the input graph looks like (i.e. what vertex/edge attributes there are, what type of data they hold etc).
Thank you for such a quick response! I only encountered a measurable leak on a graph with ~200 000 vertices. When I was running the program on smaller graphs (up to 10 000 vertices) I did not have much trouble there. I think the memory leak is relatively small. It took me quite some time to pin it down.
Attributes you used are pretty similar to mine, the name is a short string (up to 10 characters), the min_count is a list of integers and orig_id is a saved index from the unclustered graph. I am not holding any other vertex nor edge attributes. Number of edges is linear to the number of vertices (|E| <= 4|V|).
The only difference I can spot is that you are directly overwriting the variable, and I was running the computation in a separate method, leaving it for the python garbage collector.
Sorry, still no luck -- I have now tried on a graph with 200K vertices and 800K edges, and the memory usage seems to fluctuate around 180MB, sometimes a bit more, sometimes a bit less. I have changed the test code like this now:
from igraph import Graph
from random import shuffle
num_nodes = 200000
numbers = list(range(num_nodes))
shuffle(numbers)
graph = Graph.GRG(num_nodes, 0.0035)
print(graph.vcount(), graph.ecount())
graph.vs["name"] = ["name{}".format(i) for i in range(num_nodes)]
graph.vs["min_count"] = numbers
def compute(g):
strongly_connected = g.clusters(mode="strong")
condensed = strongly_connected.cluster_graph(combine_vertices=dict(name=lambda x: x, min_count='min', orig_id_graph=lambda x: x, ))
return condensed
for i in range(200000):
x = compute(graph)
Can you try to create a short, self-contained example code that reproduces the problem on your machine? Also, which OS are you using? There might be subtle differences in the memory management routines of Python on different platforms.
I am using Ubuntu 20.04. The code:
def process(G):
G.vs["orig_id_graph"] = [x.index for x in G.vs]
scc = G.clusters(mode="strong")
sizes = np.array(scc.sizes())
big_scc = set(np.where(sizes > 1)[0])
condensed = scc.cluster_graph(
combine_vertices=dict(
name=lambda x: x,
min_count='min',
orig_id_graph=lambda x: x,
)
)
singles_subgraph = condensed.induced_subgraph([x for x in condensed.vs if x.index not in big_scc])
paths = singles_subgraph.get_all_shortest_paths(0) # this is a simplification
return paths
I really could not explain the leak any other way than by the lambda function, but since you cannot replicate it it is possible I missed something else... Thank you for your efforts.
I'm kinda stuck on this as I still haven't found a clear way to reproduce this on my machine yet. Moving to the next milestone.
This issue has been automatically marked as stale because it has not had recent activity. It will be closed in 14 days if no further activity occurs. Thank you for your contributions.
@Caeph Can you test this with the latest igraph (0.9.10) and see if the problem is still there? IIRC 0.9.10 fixed some memory leaks which may have affected vertex combiners.