substructure search result optimisation
Few comments on the substructure search (ss):
-
ss should return the molecule that is the exact substructure if this molecule is in chembl eg: ss with dopamine structure does not return dopamine http://chembl-glados.herokuapp.com/g/#substructure_search_results/NCCC1%3DCC%3DC(O)C(O)%3DC1
-
the highlight switch on the left panel is hard to find the first time
-
When the highlight is on, you see the substructure in the molecules (good) but some molecules are 'tortured' (bad).
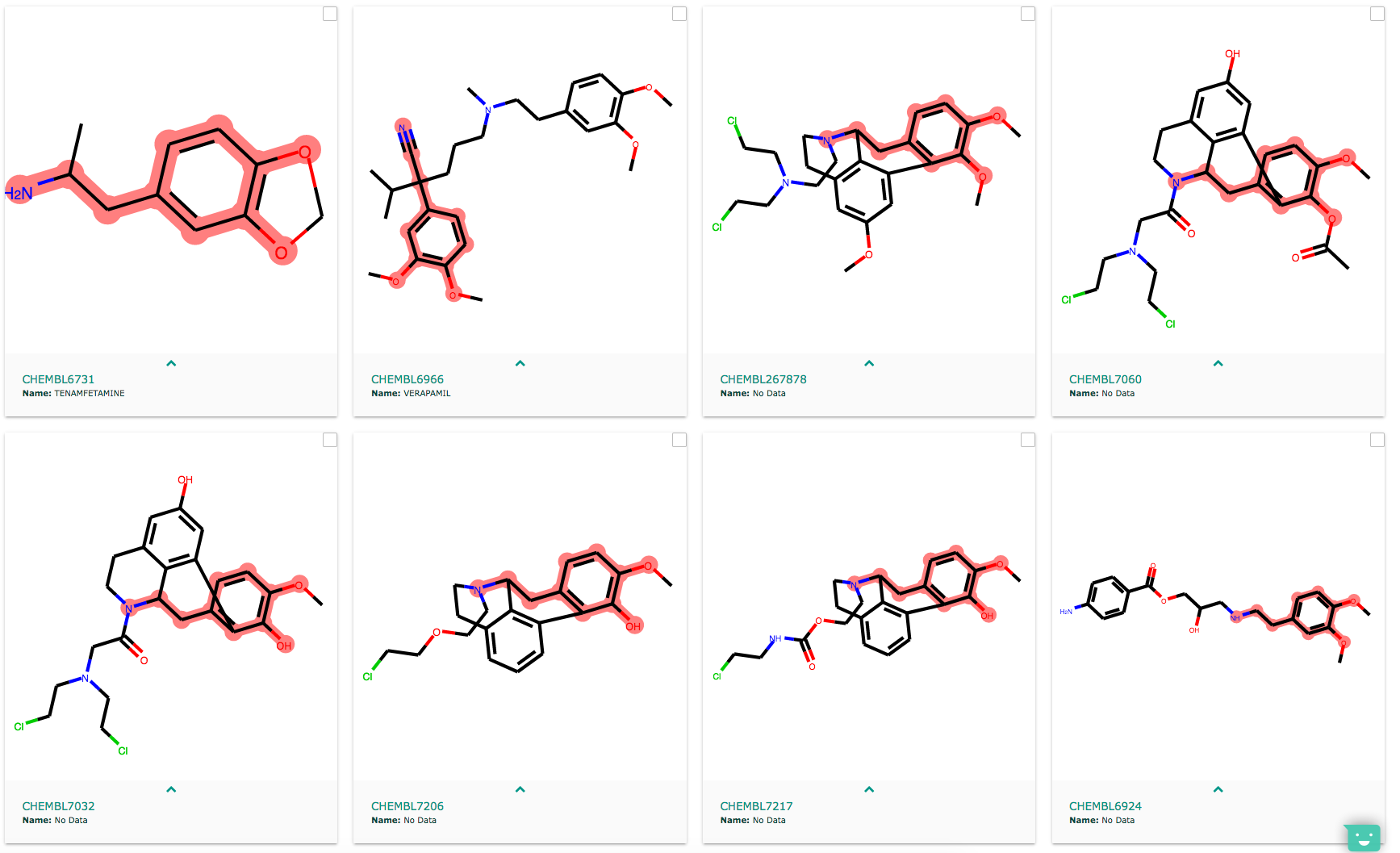
When the highlight is off, the molecules are not tortured (good), but the substructure is not highlighted (bad). Can we have highlight without molecule torture? I am very concerned about molecule well-being :)
I can't agree with the first point. Trivial results are... well, trivial and we just excluded them from similarity search as a result of @anne123 request.
I don't know yet what to do with the second point but I agree with you.
The third point is due to problems with RDKit and is already described in #499.
Point 1 is trivial if you already know your compound. In CORBEL I am working with a scientist who identified small molecules synthesised by her bacteria. These molecules are rather small so very likely to occur in other bigger molecules, so it is sensible to do a substructure search. Currently to have all the information on these compounds I need to do a substructure search + an exact match search. I mean ok, but in that case you need to say somewhere that the substructure search excludes the exact match.
Happy to further discuss this point.