Invalid k(E) at high T for BM rate rule estimate
@ChrisBNEU and I are testing my halogens pdep PR with the 1,3-hexadiene pyrolysis example (without mopac).
RMG is crashing on main branch with this error
Warning: For path reaction [CH2]C(C=CCC)C([CH2])C=CCC(50) <=> [CH2]C=CC(CC)C([CH2])C=CCC(47):
Warning: Expected kf(1896.74 K) = 2.39965e+11
Warning: Actual kf(1896.74 K) = 7.72355e+10
Warning: Expected Keq(1896.74 K) = 10.7667
Warning: Actual Keq(1896.74 K) = 0
Error: Invalid k(E) values computed for path reaction "[CH2]C(C=CCC)C([CH2])C=CCC(50) <=> [CH2]C=CC(CC)C([CH2])C=CCC(47)".
Error: Increasing number of grains did not decrease error enough (Current badness: 0.5, previous 0.5). Something must be wrong with network PDepNetwork #13
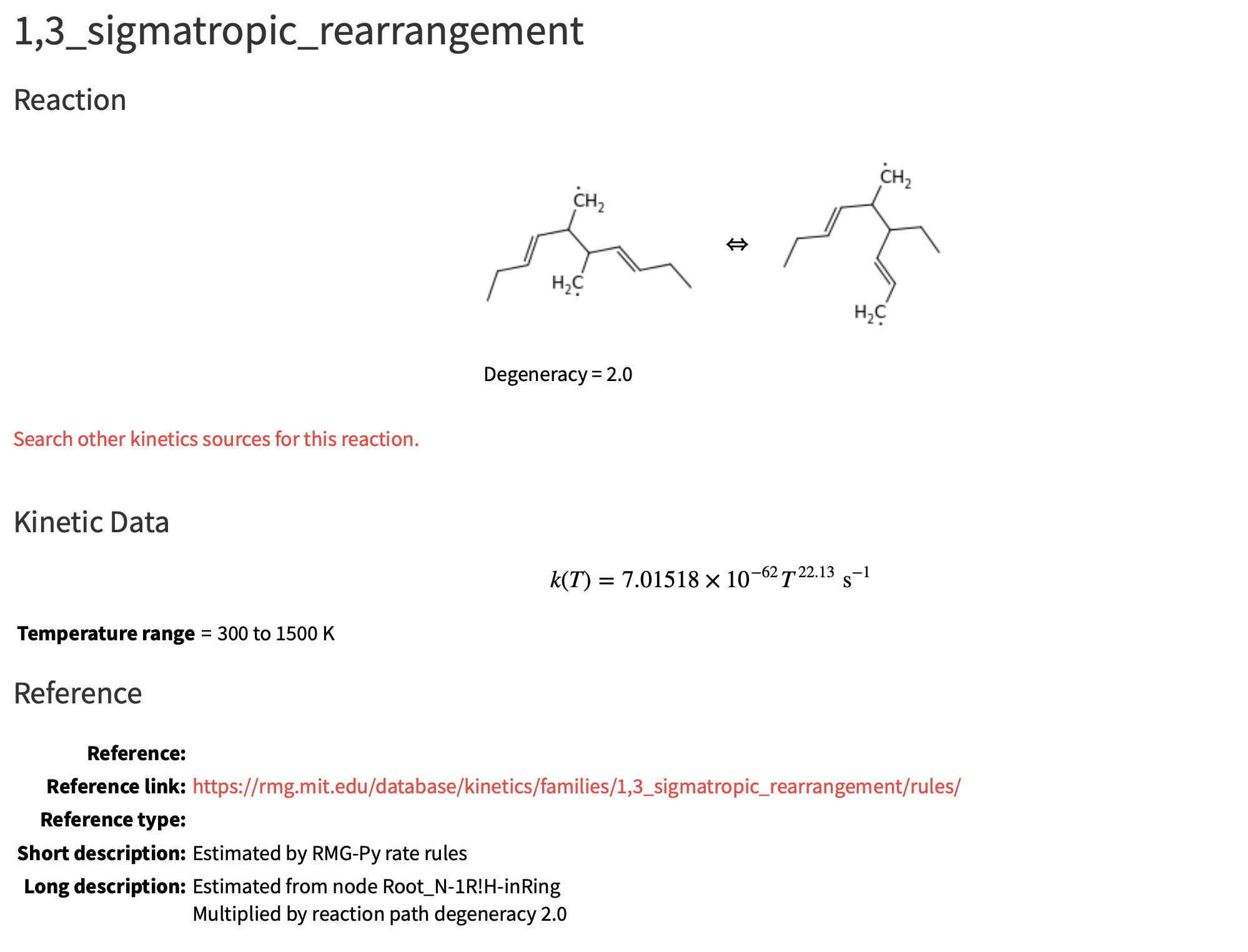
Not sure if this is due to a bad microcanonical rate due to poor density of state estimates from group freqs, or a poor high pressure rate at 1900 K (or both). Since the high P kinetics are fit to 300 - 1500 K range, and the T exponent is ~22, the rate may be too fast at T > 1500 K
Thanks @mjohnson541 for notifying me of this issue. I recently created the 1,3 sigmatropic rearrangement reaction family (PR 532 on RMG-database) that is involved in this issue and wanted to share an update.
Based on the picture above, the problem is with this node in the rate tree: Root_N-1R!H-inRing. Previously, the estimate at this node was rather interesting... (link)
label = "Root_N-1R!H-inRing",
ArrheniusBM(A=(3.50759e-62,'s^-1'), n=22.1275, w0=(750402,'J/mol'), E0=(-20429.3,'J/mol') ...
Total Standard Deviation in ln(k): 13.494121433083981"""),
However, I recently made a new PR on RMG-database (PR 550) that adds a thermo library for the species involved in the training reactions I had added that were used to fit this tree. On the new PR, the rate estimate for this specific node looks more reasonable and the standard deviation decreased (link)
label = "Root_N-1R!H-inRing",
kinetics = ArrheniusBM(A=(1.69526e+13,'s^-1'), n=-0.0541346, w0=(705700,'J/mol'), ...
Total Standard Deviation in ln(k): 8.759382282596956"""),
I do not know how relevant this change in the rate estimate is vs. other potential issues, perhaps RMG struggling with biradicals? None of the training reactions I had added involve radicals (let alone biradicals) so perhaps the thermo I added for the stable species will not matter for this specific reaction. Anyways, just wanted to share this info
This issue is being automatically marked as stale because it has not received any interaction in the last 90 days. Please leave a comment if this is still a relevant issue, otherwise it will automatically be closed in 30 days.