trying to match tf.nn.conv2d vs keras2c
I've been having issues matching the output of conv2d from keras to the output of keras2c. I've been trying to unit test the k2c_conv2d vs the output of tf.nn.conv2d.
Below is the code that I used to generate my "truth" using tf.nn.conv2d. I have an 80x80 chip of ones convolved with a 7x7 kernel of ones. The script test_conv2d.py produces two outputs. The first output is the result of the convolution using a stride of 1 and the second output is the result of the convolution using a stride of 2. In the first case, the output of the first 3 rows is not 49. This makes sense since the mask goes outside the image and with padding set to 'same', the assumed value outside the image is assumed zero. In the second case, the first row and the last two rows is not 49 as expected.
# test_conv2d.py
import numpy as np
import tensorflow as tf
x_in = np.ones((1,80,80,1),dtype='float32')
kernel_in = np.ones((7,7,1,1),dtype='float32')
x = tf.constant(x_in, dtype=tf.float32)
kernel = tf.constant(kernel_in, dtype=tf.float32)
result = tf.nn.conv2d(x, kernel, strides=[1, 1], padding='SAME')
arr = result[0,:,:,0]
np.save('conv2d_example_s1.npy',arr.numpy())
result2 = tf.nn.conv2d(x, kernel, strides=[2, 2], padding='SAME')
arr2 = result2[0,:,:,0]
np.save('conv2d_example_s2.npy',arr2.numpy())
I'm assuming that in order to match with keras2c, I need to pad the image to an 86x86 (by adding 3 rows and columns on each side set to zero), then convolve the padded image to generate an output image of 80x80 for stride 1 and 40x40 for stride 2.
Below is the C code that I used to perform the convolution in k2c.
/* test2d.c */
#include "k2c_include.h"
#include <stdio.h>
#define num_channels 2
float input_buffer[80*80] = {0};
k2c_tensor input = {&input_buffer[0],num_channels,80*80,{ 80,80, 1, 1, 1}};
float pad_buffer[86*86]= {0};
k2c_tensor pad = {&pad_buffer[0],num_channels,86*86,{86,86,1,1,1}};
float output_buffer[86*86] = {0};
k2c_tensor output = {&output_buffer[0],num_channels,80*80,{ 80,80, 1, 1, 1}};
k2c_tensor output2 = {&output_buffer[0],num_channels,40*40,{ 40,40, 1, 1, 1}};
float kernel_buffer[7*7] = {0};
k2c_tensor kernel = {&kernel_buffer[0],num_channels,7*7,{ 7,7, 1, 1, 1}};
float bias_buffer = 0;
k2c_tensor bias = {&bias_buffer,num_channels,1,{ 1,1, 1, 1, 1}};
size_t stride[num_channels] = {1,1};
size_t dilation[num_channels] = {0};
size_t pad_values[4] = {3,3,3,3};
void print_tensor(k2c_tensor* output)
{
int k=0;
for (int i=0;i<output->shape[0];++i) {
for (int j=0;j<output->shape[1];++j,++k) {
printf("%.9f ",output->array[k]);
}
printf("\n");
}
}
int main(int argc, char** argv)
{
if (argc >= 3) {
stride[0] = atoi(argv[1]);
stride[1] = atoi(argv[2]);
}
for (int i=0;i<80*80;++i) {
input_buffer[i] = 1;
}
for (int i=0;i<7*7;++i) {
kernel_buffer[i] = 1;
}
/**
* 2D (spatial) Padding.
*
* :param output: tensor to store padded output data.
* :param input: tensor to pad.
* :param fill: value to fill in padded areas.
* :param pad: array[4] of how many rows/cols to pad. Order is {before dim 1, after dim 1, before dim 2, after dim 2}.
*/
k2c_pad2d(&pad, &input, 0.0f, pad_values);
// print_tensor(&pad);
//return 0;
if (stride[0] == 1) {
k2c_conv2d(&output, &pad, &kernel,
&bias, stride, dilation,
k2c_linear);
print_tensor(&output);
} else {
k2c_conv2d(&output2, &pad, &kernel,
&bias, stride, dilation,
k2c_linear);
print_tensor(&output2);
}
return 0;
}
I generated the resulting chips using these commands:
./test2d.exe 1 1 > d1.txt && ./test2d.exe 2 2 > d2.txt
Then used the following script to compare the output from tensorflow vs k2c:
# show_pair.py
import numpy as np
import matplotlib.pyplot as plt
def show_pair():
d1 = np.loadtxt('d1.txt')
d2 = np.loadtxt('d2.txt')
p1 = np.load('conv2d_example_s1.npy')
p2 = np.load('conv2d_example_s2.npy')
plt.figure()
plt.subplot(2,2,1); plt.matshow(d1,fignum=False); plt.title('k2c stride 1')
plt.subplot(2,2,2); plt.matshow(p1,fignum=False); plt.title('tf stride 1')
plt.subplot(2,2,3); plt.matshow(d2,fignum=False); plt.title('k2c stride 2')
plt.subplot(2,2,4); plt.matshow(p2,fignum=False); plt.title('tf stride 2')
plt.show()
if __name__ == '__main__':
show_pair()
Attached is the image that I generated.
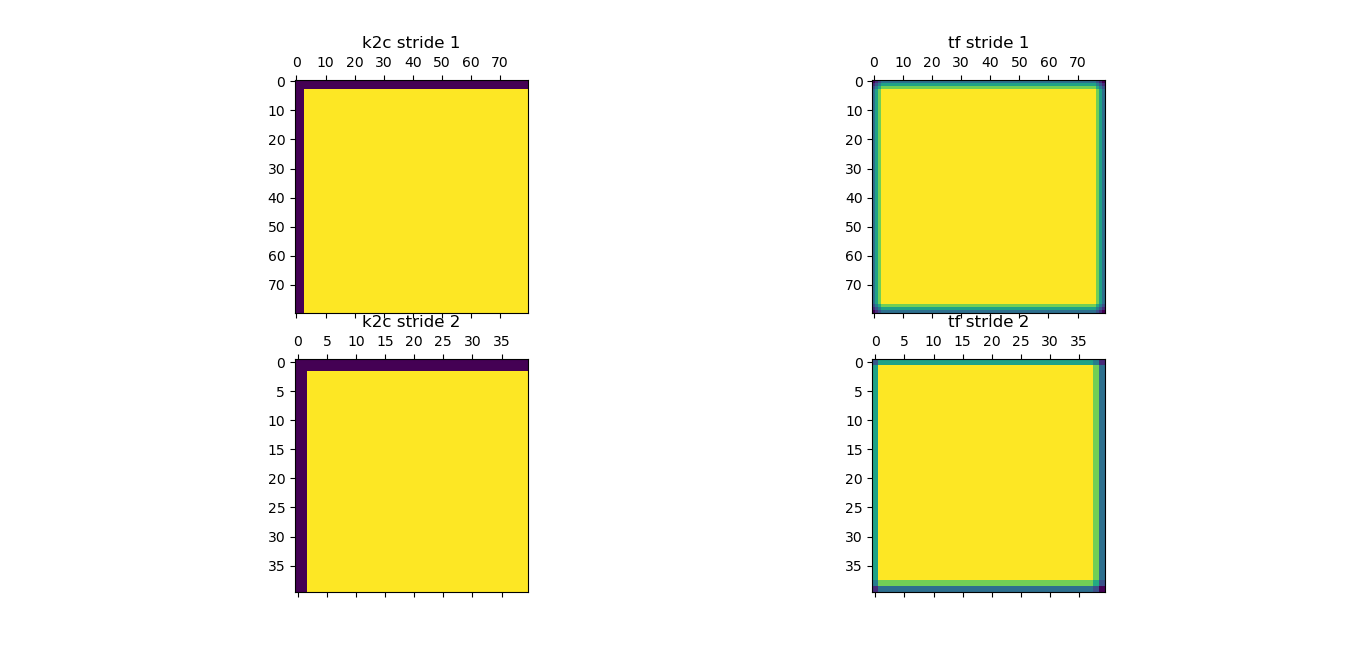
First, notice there is not gradient in the output of k2c. Second, pixel (2,2) is the first value on the k2c convolution with stride 2 with value 49 vs pixel(1,1) from the output of tf with stride 2.
I maybe simulating the tf.nn.conv2d incorrectly in C. Feedback in how to do it correctly is welcomed. But I think the shift of the stride is still different.
Your help will be greatly appreciated
The output was generated using conda with tensorflow cpu 2.3.0 vs the master branch of k2c on windows.
The C code above had dilation=0. When I run it with dilation=1, I get closer results but there is a different shift when the stride is 2.

I was able to match the k2c conv2d with keras. The trick is to specify the padding and the size of the padded image according to the rules in https://www.tensorflow.org/api_docs/python/tf/nn#notes_on_padding_2. The code below has the modification:
#include "k2c_include.h"
#include <stdio.h>
#include <math.h>
#include <algorithm>
#define num_channels 2
float input_buffer[80*80] = {0};
k2c_tensor input = {&input_buffer[0],num_channels,80*80,{ 80,80, 1, 1, 1}};
float pad_buffer[86*86]= {0};
k2c_tensor pad = {&pad_buffer[0],num_channels,86*86,{86,86,1,1,1}};
float output_buffer[86*86] = {0};
k2c_tensor output = {&output_buffer[0],num_channels,80*80,{ 80,80, 1, 1, 1}};
k2c_tensor output2 = {&output_buffer[0],num_channels,40*40,{ 40,40, 1, 1, 1}};
float kernel_buffer[7*7] = {0};
k2c_tensor kernel = {&kernel_buffer[0],num_channels,7*7,{ 7,7, 1, 1, 1}};
float bias_buffer = 0;
k2c_tensor bias = {&bias_buffer,num_channels,1,{ 1,1, 1, 1, 1}};
size_t stride[num_channels] = {1,1};
size_t dilation[num_channels] = {1,1};
size_t pad_values[4] = {3,3,3,3};
void print_tensor(k2c_tensor* output)
{
int k=0;
for (int i=0;i<output->shape[0];++i) {
for (int j=0;j<output->shape[1];++j,++k) {
printf("%.9f ",output->array[k]);
}
printf("\n");
}
}
int main(int argc, char** argv)
{
if (argc >= 3) {
stride[0] = atoi(argv[1]);
stride[1] = atoi(argv[2]);
}
for (int i=0;i<80*80;++i) {
input_buffer[i] = 1;
}
for (int i=0;i<7*7;++i) {
kernel_buffer[i] = 1;
}
/**
* 2D (spatial) Padding.
*
* :param output: tensor to store padded output data.
* :param input: tensor to pad.
* :param fill: value to fill in padded areas.
* :param pad: array[4] of how many rows/cols to pad. Order is {before dim 1, after dim 1, before dim 2, after dim 2}.
*/
int in_height = 80;
int in_width = 80;
int filter_height = 7;
int filter_width = 7;
int stride_height = stride[0];
int stride_width = stride[1];
int pad_along_height = 0;
int pad_along_width = 0;
if ( (in_height % stride_height) == 0) {
pad_along_height = std::max(filter_height - stride_height, 0);
} else {
pad_along_height = std::max(filter_height - (in_height % stride_height), 0);
}
if ( (in_width % stride_width) == 0) {
pad_along_width = std::max(filter_width - stride_width, 0);
} else {
pad_along_width = std::max(filter_width - (in_width % stride_width), 0);
}
int pad_top = pad_along_height / 2;
int pad_bottom = pad_along_height - pad_top;
int pad_left = pad_along_width / 2;
int pad_right = pad_along_width - pad_left;
pad_values[0] = pad_top;
pad_values[1] = pad_bottom;
pad_values[2] = pad_left;
pad_values[3] = pad_right;
pad.numel = (in_height+pad_top+pad_bottom)
*(in_width+pad_left+pad_right);
pad.shape[0] = in_height+pad_top+pad_bottom;
pad.shape[1] = in_width+pad_left+pad_right;
k2c_pad2d(&pad, &input, 0.0f, pad_values);
// print_tensor(&pad);
//return 0;
if (stride[0] == 1) {
k2c_conv2d(&output, &pad, &kernel,
&bias, stride, dilation,
k2c_linear);
print_tensor(&output);
} else {
k2c_conv2d(&output2, &pad, &kernel,
&bias, stride, dilation,
k2c_linear);
print_tensor(&output2);
}
return 0;
}
And the resulting images look as follows:

The code in weights2c.py seems to have the computations for the padding. But they don't look the same as the notes in the link above.