Erronous segmentation of the whole stack, But a cropped region works perfectly.
I am having this issue with cellpose unable to segment nuclei when I provide with the whole stack. But it does work fine with a small cropped region of the same image. Why is this so?
The voxel size of cropped and uncropped images is the same, only the pixels are cropped in the XYZ directions.
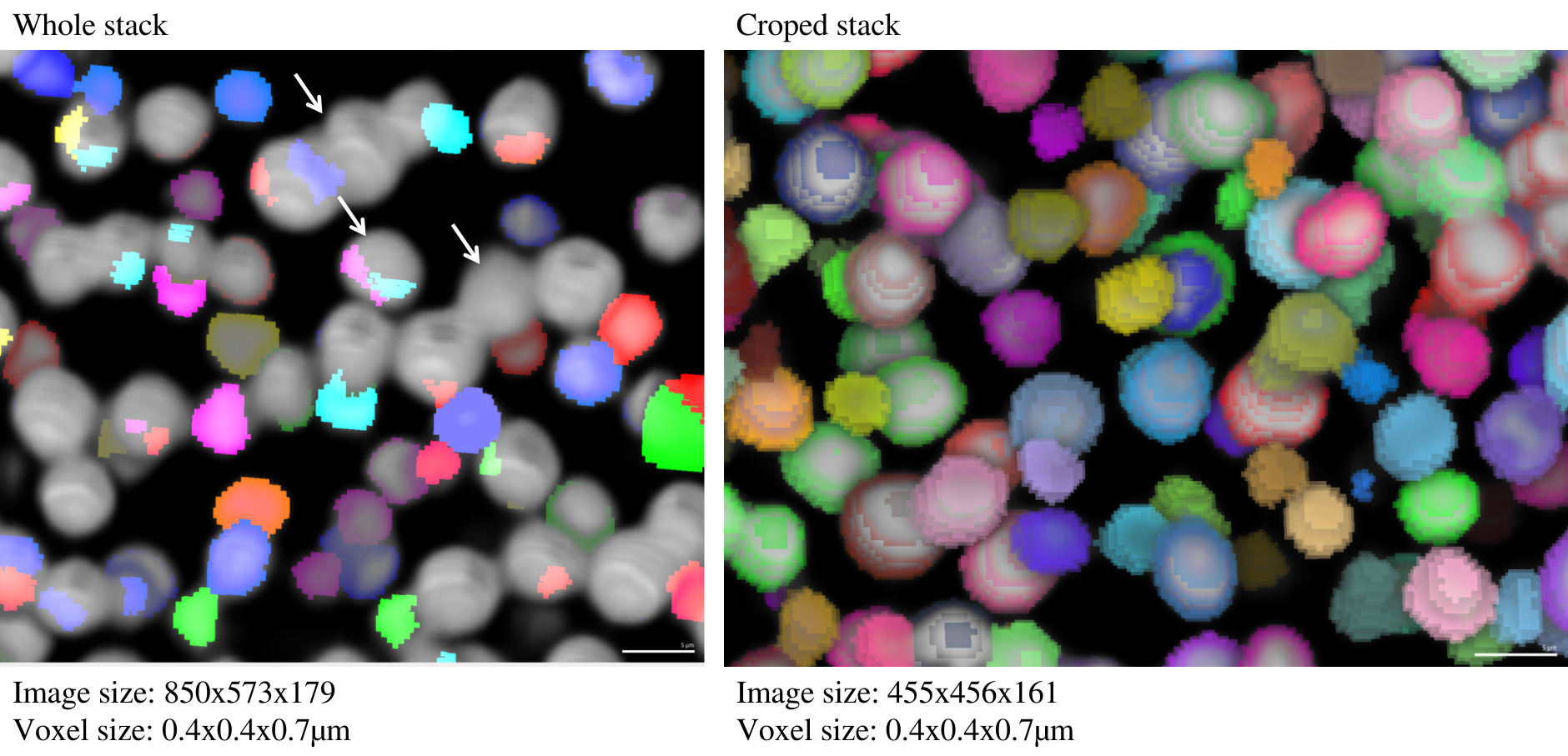

@athulrv, could you provide the code you used to get your results? Given the blockiness of the segmentation, I'm guessing there was some pretty dramatic rescaling of the image, and it's possible that this issue has to do with how that rescaling (maybe automatic scaling using the size model?) changes depending on the crop size.
I'm also curious what tool you are using for the visualization of those 3D labels, that looks nice.
Hi @kevinjohncutler I have just used the cellpose GUI for performing the segmentation, not the codes.
I am not sure what the Scale disk On and Off does? I already tried to change the cell diameter in this case. I wouldn't mind sharing the raw image used here.
The software used for 3D visualization is "MorphographX" open source- https://morphographx.org/
@athulrv, thanks for sharing that link. The scale disk is only for visualizing cell diameters, if I recall correctly. It might be useful for you to share screenshots of the GUI. Also feel free to share the file here and I will do some investigation.
Hi @kevinjohncutler I rerun this with the command line cellpose- "python -m cellpose --dir ~/Cellpose_test/ --pretrained_model nuclei --diameter 15. --save_tif --use_gpu --do_3D"
I have also run the same with diameter 0. The results are not optimal compared to the cropped input stack.
Image size; Voxel size 1024x1024x267 0.399796x0.399796x0.699998µm
Please find the image I am trying to segment from here- "Uncropped_input.tif" https://nextcloud.mpipz.mpg.de/s/AntbK5YsQxR8Gd6 You will also find the cropped stack which was near perfectly segmented by cellpose.
Thanks a lot in advance
Best, Athul
@athulrv, thanks for sharing your data. I took a look and I think I found your problem. Your cropped image is more densely populated than the full image. If you take a look at the output of transforms.normalize99() on your volumes, the cropped one retains cell features, but the full one gets blown out:
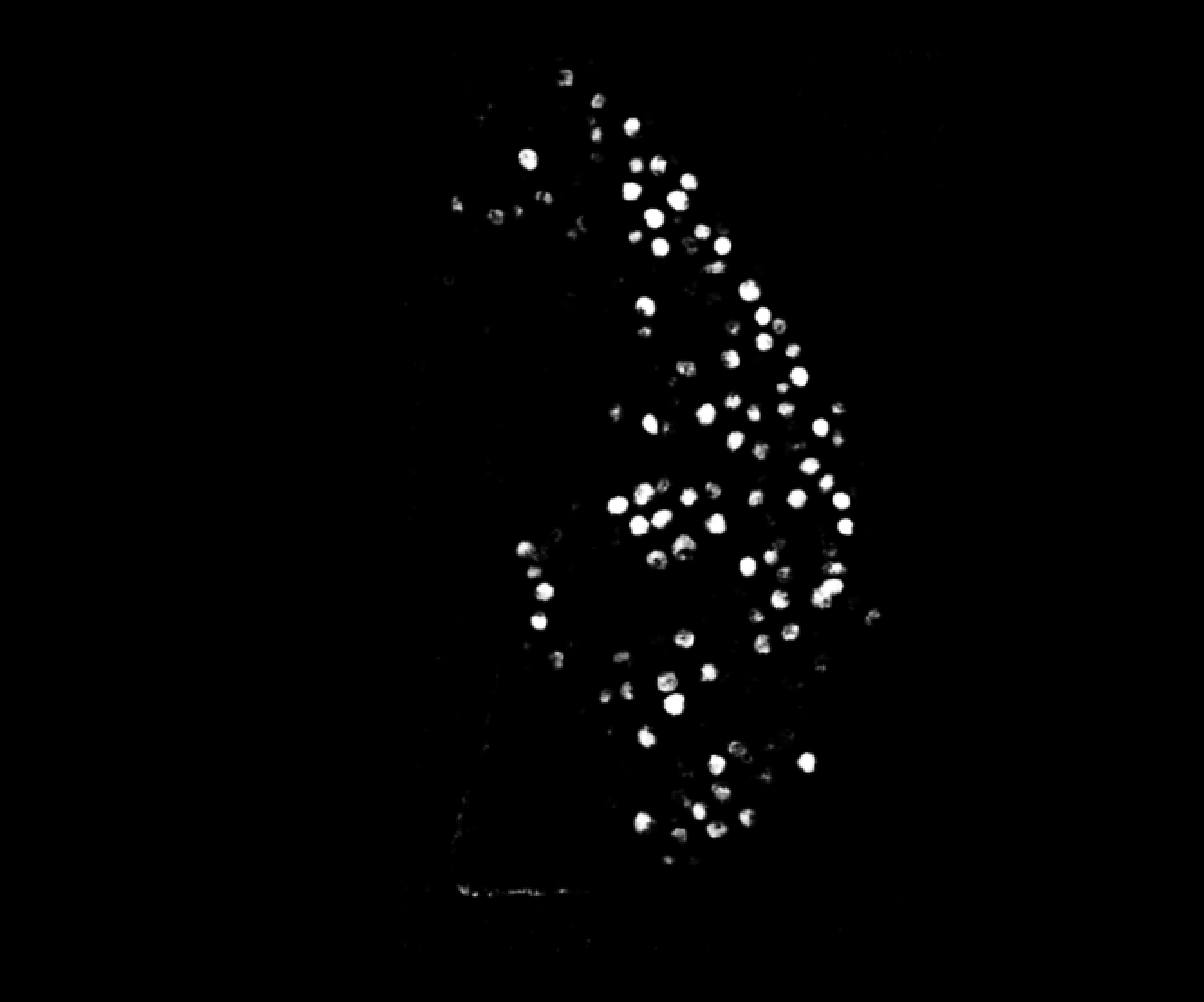
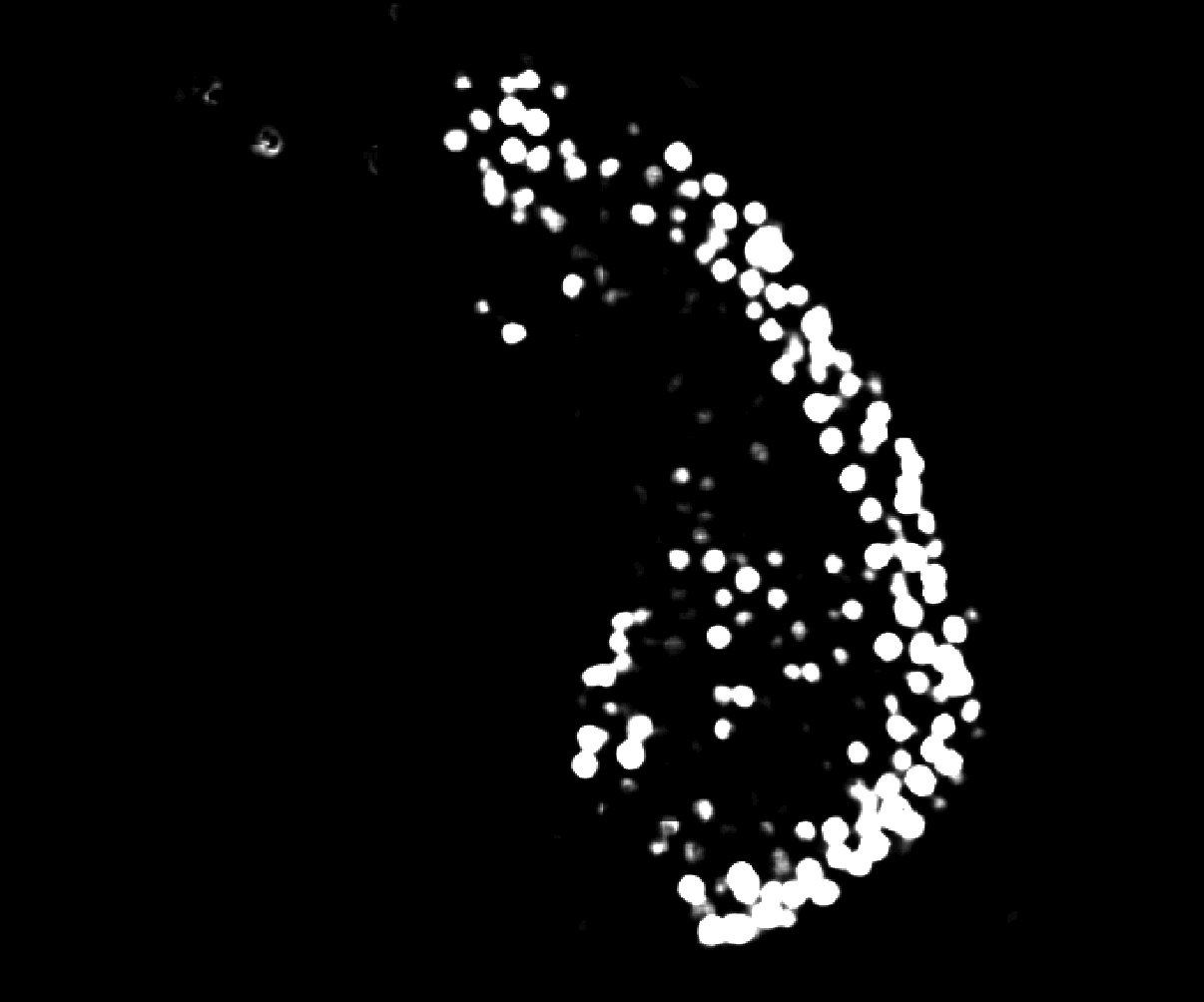
The effect of this is that there is no contrast between cells and they cannot be segmented. The reason this happens in normalize99 is that the default percentile range is 1 to 99, but that crushes the image features when most pixels are in the background and the bright cell pixels are in the top 1% (so cell pixels all get set to 1). I encountered this bug myself a long while ago with bacterial time lapses, where at the start of the movie you have a very sparse field of cells. The fix is simple and I included it in Omnipose: change the range to 0.01 and 99.99. This excludes outliers (as is the intent) but without the crushing effect we observe in your data.
However, I need to publish a version of Omnipose with the 3D bugs ironed out. I suggest that, in the meantime, you just edit the normalize99 function in your own copy of Cellpose to the range I suggested (possibly even 0.001 and 99.999 for 3D volumes).
@kevinjohncutler. Thanks a lot for the update.
Just that it's clear for me. How/Where do I change the "normalize99" parameter? Is it possible when using gui?
Best, Athul
@athulrv Normalize99 is not a parameter, but a function. You can correct its behavior by using the Omnipose version. You can try setting omni=True in the GUI (I think there should be a button for that still). Not sure if that will affect the normalization before running the network, but I think it should. But otherwise you'll need to make sure you've installed omnipose with pip install omnipose, download cellpose from github, use pip install -e <path to cellpose> to install it, and then edit the transforms.py file and change omni=False to omni=True. Alternately, instead of installing Omnipose, you could just the whole function in cellpose with mine in omnipose.utils:
def normalize99(Y,lower=0.01,upper=99.99):
""" normalize image so 0.0 is 0.01st percentile and 1.0 is 99.99th percentile """
X = Y.copy()
return np.interp(X, (np.percentile(X, lower), np.percentile(X, upper)), (0, 1))
Thanks a lot @kevinjohncutler, That worked for me.
Best, Athul
Great! @carsen-stringer you may want to consider changing normalize99 defaults in a future release.