Running simulation on HPCC
I've run a few simulations on my laptop and everything works fine. Now I am trying to use the HPCC at my university to test the simulation, but I encountered this error while viewing the output images.
I have no idea what this error means exactly. I also notice that for the first few minutes of the simulation, there is nothing wrong with the images, however, this error message comes up in the middle of the simulation after a while. In the meantime, this error doesn't stop my simulation, instead, it keeps running till the end but the rest of the images are all having the same issue.
We would definitely need more information about your situation to even begin to try to help you debug it. From your error, I'm guessing your .svg files have been corrupted - either they weren't completely written/saved or there's a memory error or something else. You'd want to be sure you haven't run up against limits on your HPC account, like storage limits for your output files or runtime limits, although you claim that your simulation continues to run. A few suggestions/questions:
- if you're willing to share your model, we could try to run it to see if by chance we could reproduce the problem
- you're running with identical parameters, i.e., the very same .xml configuration file, on both your laptop and the HPC?
- just to confirm, you're not trying to run it as a distributed memory program, e.g., using MPI, on the HPC are you?
- do you have any insights into how the HPC does file I/O? Would it be possible that output files are being written onto some unusual disk system where such I/O problems have appeared for others (probably unlikely)
- have you attempted to describe your problem to the HPC support staff? They may have some insights and suggestions.
cancer_immune.zip coor_anti.txt I have included both my codes and the data I used here. To answer your questions
- I am running with identical parameters on both my laptop and the HPC.
- I'm not trying to run it as a distributed memory program.
- I don't have any insights into how the HPC does file I/O. But I can double-check both this question and the issue of running up against limits with the HPCC tech support from my university.
- I have described this problem to the HPC tech support, but they had no idea why this error message popped up.
What release of PhysiCell are you using - the latest, 1.11.0 ? (see VERSION.txt in your root directory)
And did you attempt to run the original cancer_immune_3D model on the HPC machine? Just curious about a possible link between the error that occurred and the edits you made.
I assume when you run it on the HPC, you edit the path in custom_modules/cancer_immune_3D.cpp when opening coor_anti.txt, e.g., when I ran it on my laptop, I just put that file in the root directory and change the code to coordinates.open("coor_anti.txt", std::ios::in);
Here's a snapshot from the simulation running on my laptop. Keep in mind the .svg might be a bit strange for a 3D model, as it's only capturing cells that intersect the Z=0 plane.
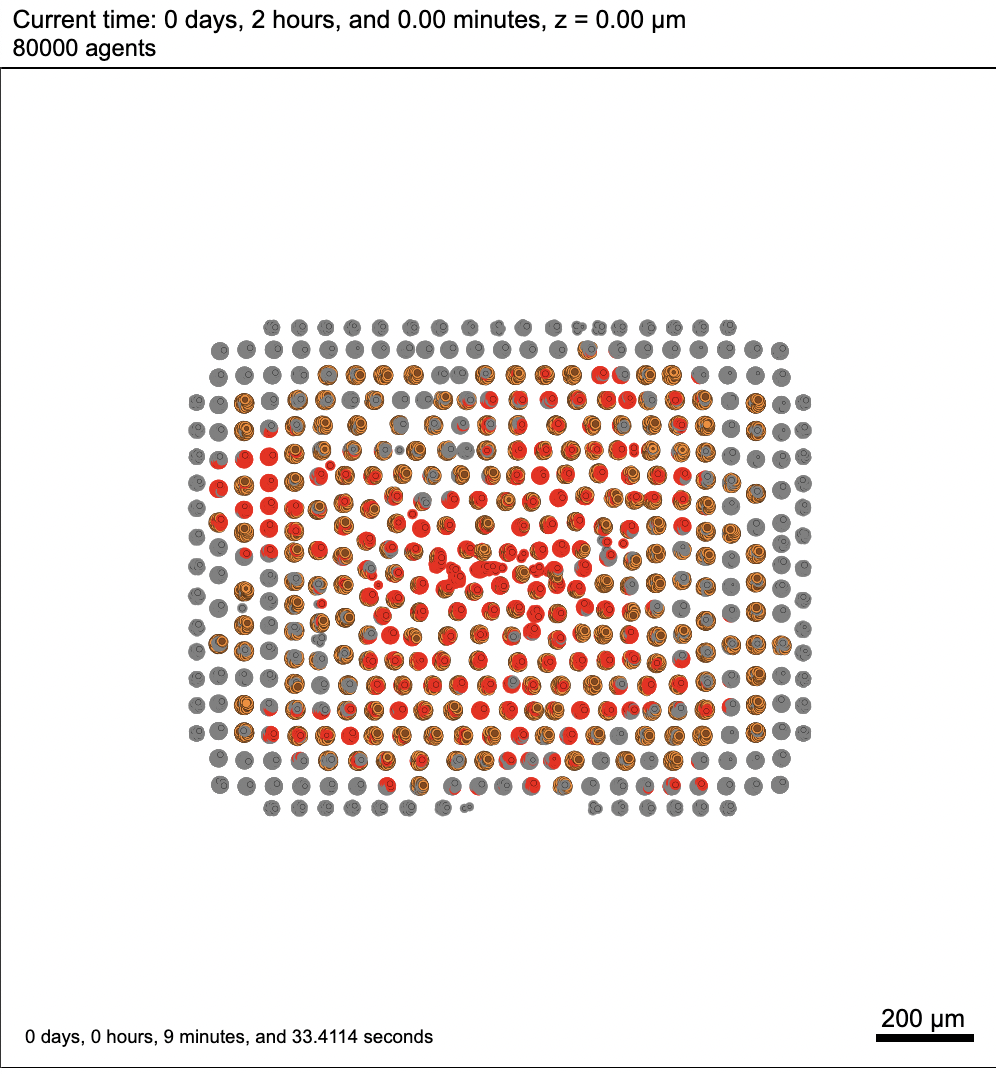

The version I'm using is 1.9.1.
I didn't try the original cancer immune 3D on HPCC. What you have is the same as the ones I have in the beginning. The error message shows up after about 30 images (or even later).
For starters, I'd strongly recommend updating to the latest PhysiCell release. I created a repo where I did that, using your code and
- updated the Makefile for 1.11
- inserted some additional sanity checks in the setup_tissue(), e.g., to show me the range of cell positions (do these seems correct?)
- edited your .xml where you seemed to have open-ended comments in <user_parameters> (however, the code still seems to parse the xml correctly)
- included a rwh1.xml and /rwh1, for my own testing, but you can ignore
- there are more model params in 1.11, but they should take on safe default values and therefore I didn't include them in the .xml
- yes, there are some risks in taking an older .xml model and using in a newer release, but hopefully you're at least willing to try it
That repo is at https://github.com/rheiland/tony_model . I can let this model run longer on my laptop at some point to see if I get the mangled .svg you reported. But it would be good if you could try it on your HPC system too.
Out of curiosity, is there some reason you are running a 3D model, as opposed to a 2D version? I'm just thinking the initial debugging and results would go smoother. Then switch to 3D once you're confident in the 2D results?
Just pushed an updated main.cpp that uses the _v2 version of saving full data.
Here's a .svg well into the simulation. But this is running on my Mac. It's hard to imagine there's a fundamental problem running the same sim on your HPC machine.
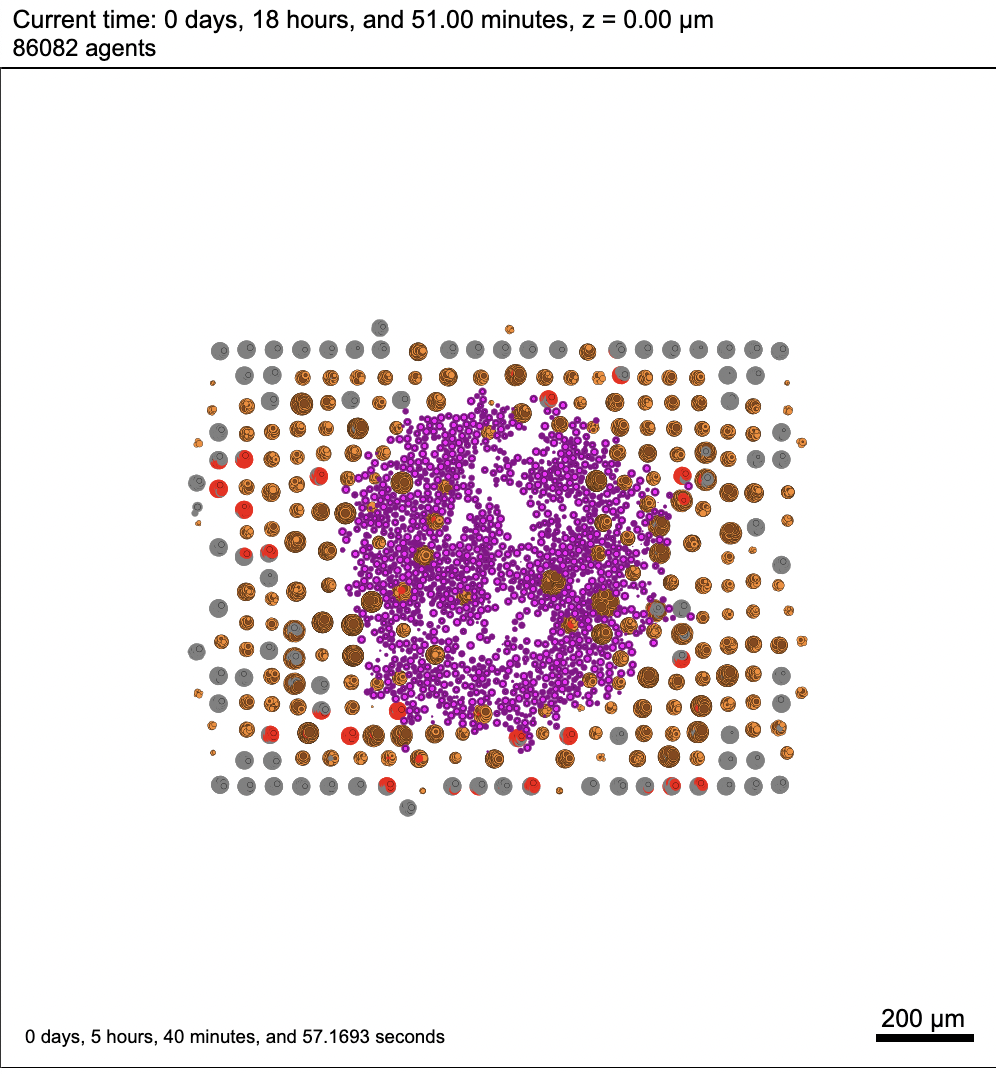

Here are some renderings using the fairly primitive 3D vis available in PhysiCell Studio. Currently it only colors cells by their cell type ID. It seems you have spheroids of cells?
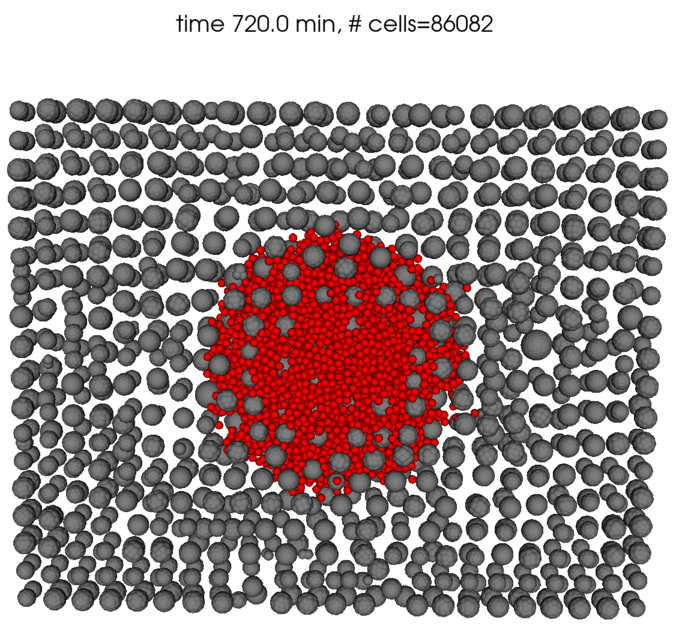
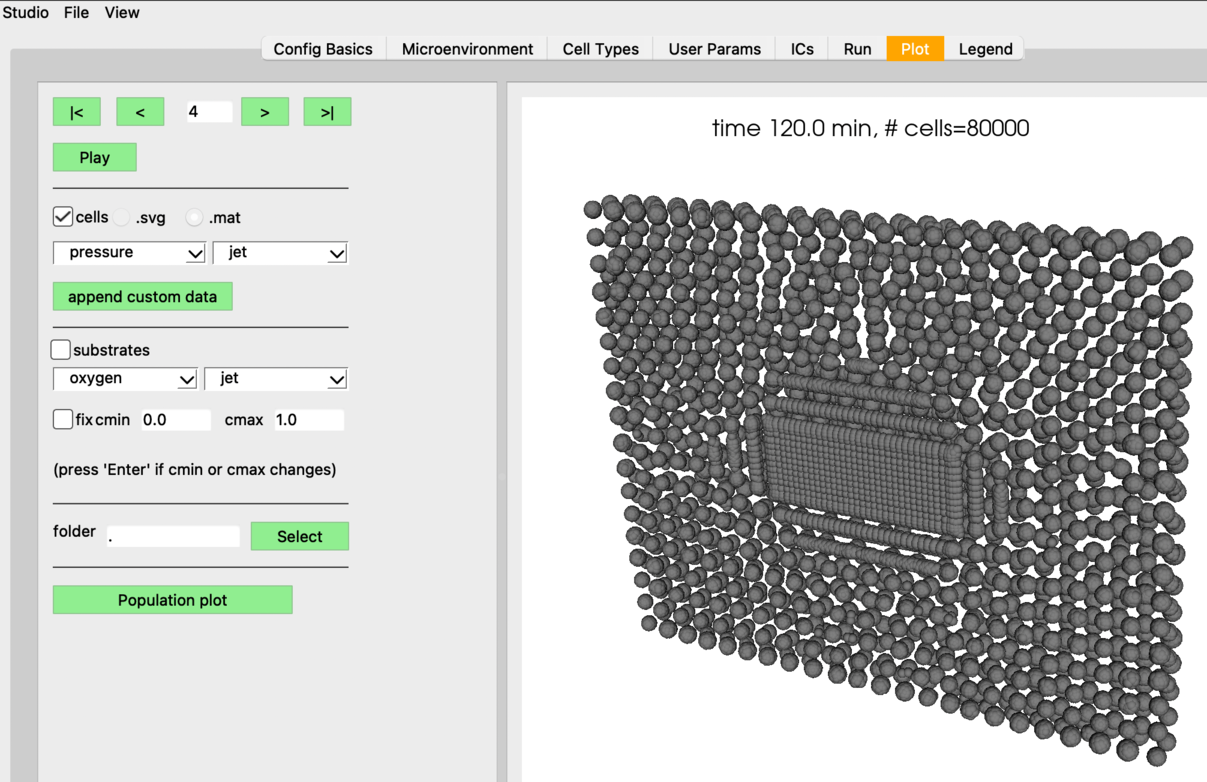

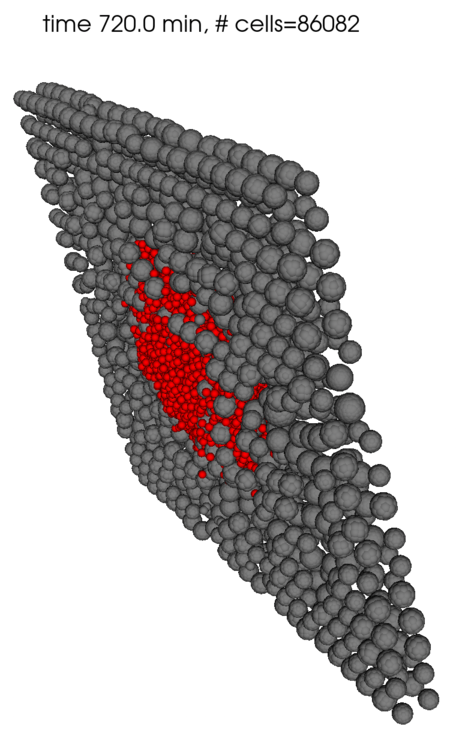
I will try to use the latest version on HPCC and see if that makes any difference.
As for the model, our project requires to build a 3D instead of a 2D model, so I chose to start with 3D model.
For the cells' positions, I used the given data to assign positions for every cancer cell initially. In this case, red cells denote the cancer cells while grey cells denote healthy cells (tissue). After a certain period of time, I introduced immune cells (colored in purple) in the middle as a spheroid.
By the way, I believe the parameters that I had in this setting are incorrect. In general, every cell should be adjacent to each other in the beginning instead of splitting up into several neighborhoods. I should have used a smaller number for cell to cell adhesion strength and bigger number for cell to cell repulsion strength.
I will talk to the HPCC tech support again with your updates, and I will follow up if the problem is still unsolved.
time to close?
Sure, please go ahead.
On Tue, Oct 1, 2024 at 10:29 AM Daniel Bergman @.***> wrote:
time to close?
— Reply to this email directly, view it on GitHub https://github.com/MathCancer/PhysiCell/issues/150#issuecomment-2386573766, or unsubscribe https://github.com/notifications/unsubscribe-auth/APNKAWRGGL34LAN4AV2STK3ZZLLYJAVCNFSM6AAAAABPGB2LS6VHI2DSMVQWIX3LMV43OSLTON2WKQ3PNVWWK3TUHMZDGOBWGU3TGNZWGY . You are receiving this because you authored the thread.Message ID: @.***>